第 111 章 Prospensity Score 傾向性評分
111.1 Practical05 - causal inference
數據還是和前一章節一樣的數據。傾向性評分的 R 包有很多,下面用 R 來進行大多數的計算。
111.1.1 初步熟悉數據內容
##
## . use "../backupfiles/RFAcat.dt. describe
##
## Contains data from ../backupfiles/RFAcat.dta
## obs: 3,551
## vars: 18 5 Nov 2013 15:05
## -------------------------------------------------------------------------------
## storage display value
## variable name type format label variable label
## -------------------------------------------------------------------------------
## id float %9.0g Patient ID
## age float %9.0g
## gender float %9.0g gender
## smoke float %9.0g smoke Smoking status
## hospital float %9.0g Hospital ID
## nodules float %9.0g Number of nodules
## mets float %9.0g Number of other metastatic sites
## duration float %9.0g Duration of disease (in months)
## maxdia float %9.0g Diameter of largest nodule (in
## cm)
## primary float %22.0g primary Location of primary cancer
## position float %9.0g position Ease with which nodules can be
## reached
## coag float %9.0g coag Coagulopathy
## rfa float %23.0g rfa Treatment variable: RFA or
## standard surgery
## dodp float %9.0g dodp Outcome variable: death or
## disease progression within 36
## months
## agecat float %9.0g agecat Age in categories
## nodcat float %9.0g nodcat Number of nodules in categories
## durcat float %12.0g durcat Duration of disease in categories
## diacat float %9.0g diacat Diameter of largest nodule in
## categories
## -------------------------------------------------------------------------------
## Sorted by: id
##
## .
## . **********************
##
## . * Explore the data *
##
## . **********************
##
## .
## . /* Question 1 */
## / is not a valid command name
## r(199);
##
## . * Exposure and outcome
##
## . tab rfa
##
## Treatment variable: RFA |
## or standard surgery | Freq. Percent Cum.
## ------------------------+-----------------------------------
## standard surgery | 1,848 52.04 52.04
## radiofrequency ablation | 1,703 47.96 100.00
## ------------------------+-----------------------------------
## Total | 3,551 100.00
##
## . tab dodp rfa, col
##
## +-------------------+
## | Key |
## |-------------------|
## | frequency |
## | column percentage |
## +-------------------+
##
## Outcome |
## variable: |
## death or |
## disease | Treatment variable:
## progressio | RFA or standard
## n within | surgery
## 36 months | standard radiofreq | Total
## -----------+----------------------+----------
## no | 1,255 1,349 | 2,604
## | 67.91 79.21 | 73.33
## -----------+----------------------+----------
## yes | 593 354 | 947
## | 32.09 20.79 | 26.67
## -----------+----------------------+----------
## Total | 1,848 1,703 | 3,551
## | 100.00 100.00 | 100.00
##
## .
## . * New (categorised) variables:
##
## . tab1 agecat nodcat durcat diacat, m
##
## -> tabulation of agecat
##
## Age in |
## categories | Freq. Percent Cum.
## ------------+-----------------------------------
## <45 | 96 2.70 2.70
## 45-49 | 860 24.22 26.92
## 50-54 | 1,755 49.42 76.34
## 55-59 | 662 18.64 94.99
## 60-64 | 135 3.80 98.79
## 65+ | 43 1.21 100.00
## ------------+-----------------------------------
## Total | 3,551 100.00
##
## -> tabulation of nodcat
##
## Number of |
## nodules in |
## categories | Freq. Percent Cum.
## ------------+-----------------------------------
## 1 | 997 28.08 28.08
## 2 | 1,350 38.02 66.09
## 3 | 635 17.88 83.98
## 4 | 204 5.74 89.72
## 5-9 | 304 8.56 98.28
## 10+ | 61 1.72 100.00
## ------------+-----------------------------------
## Total | 3,551 100.00
##
## -> tabulation of durcat
##
## Duration of |
## disease in |
## categories | Freq. Percent Cum.
## -------------+-----------------------------------
## <10 months | 53 1.49 1.49
## 10-19 months | 1,437 40.47 41.96
## 20-29 months | 1,528 43.03 84.99
## 30-39 months | 397 11.18 96.17
## 40+ months | 136 3.83 100.00
## -------------+-----------------------------------
## Total | 3,551 100.00
##
## -> tabulation of diacat
##
## Diameter of |
## largest |
## nodule in |
## categories | Freq. Percent Cum.
## ------------+-----------------------------------
## <1.5 cm | 967 27.23 27.23
## 1.5-1.9cm | 1,147 32.30 59.53
## 2-2.4cm | 940 26.47 86.00
## 2.5-2.9cm | 403 11.35 97.35
## 3cm+ | 94 2.65 100.00
## ------------+-----------------------------------
## Total | 3,551 100.00
##
## .
## .111.1.2 把連續型變量以分類型數據的形式放入模型中:
***************************
* Regression adjustment *
***************************
/* Question 2 */
// teffects ra (dodp i.agecat gender i.smoke i.hospital i.nodcat ///
// i.mets i.durcat i.diacat i.primary i.position, ///
// logit) (rfa)你會發現 STATA 停不下來,計算永遠都不會收斂。這是因爲我們在這個模型中結果部分加入了太多的分類型變量,但數據又沒辦法進行足夠的計算。
111.1.3 用相同的模型結構估計每個人的傾向性評分
RFAcat <- read_dta("../backupfiles/RFAcat.dta")
RFAcat <- RFAcat %>%
mutate(gender = factor(gender, labels = c("men", "women")),
smoke = factor(smoke, labels = c("never", "ex", "current")),
hospital = as.factor(hospital),
primary = factor(primary, labels = c("bladder",
"breast",
"colorectal",
"gullet",
"kidney",
"prostate",
"skin",
"stomach",
"testicular")),
position = factor(position, labels = c("easy", "moderate", "difficult")),
nodcat = factor(nodcat, labels = c("1","2","3","4","5-9","10+")),
mets = as.factor(mets),
agecat = factor(agecat, labels = c("< 45", "45-49", "50-54", "55-59", "60-64","65+")),
durcat = factor(durcat, labels = c("< 10m", "10-19m", "20-29m", "30-39m", "40+m")),
diacat = factor(diacat, labels = c("<1.5cm", "1.5-1.9cm", "2-2.4cm", "2.5-2.9cm",
"3cm+")))
Pros_Score <- glm(rfa ~ agecat + gender + smoke + hospital + nodcat + mets +
durcat + diacat + primary + position, family = binomial(link = logit),
data = RFAcat)
summary(Pros_Score)##
## Call:
## glm(formula = rfa ~ agecat + gender + smoke + hospital + nodcat +
## mets + durcat + diacat + primary + position, family = binomial(link = logit),
## data = RFAcat)
##
## Deviance Residuals:
## Min 1Q Median 3Q Max
## -2.45628 -0.90333 -0.26702 0.90927 2.67426
##
## Coefficients:
## Estimate Std. Error z value Pr(>|z|)
## (Intercept) 0.243514 0.505730 0.4815 0.630154
## agecat45-49 0.090044 0.261908 0.3438 0.730996
## agecat50-54 0.107064 0.255438 0.4191 0.675115
## agecat55-59 -0.040906 0.269042 -0.1520 0.879154
## agecat60-64 0.074712 0.327138 0.2284 0.819351
## agecat65+ 0.398326 0.431966 0.9221 0.356464
## genderwomen 0.083521 0.151596 0.5509 0.581673
## smokeex 0.117319 0.150316 0.7805 0.435105
## smokecurrent 0.144515 0.104757 1.3795 0.167732
## hospital2 -0.190892 0.140877 -1.3550 0.175407
## hospital3 0.345035 0.138004 2.5002 0.012413 *
## hospital4 -0.094451 0.191935 -0.4921 0.622648
## nodcat2 -0.085830 0.097834 -0.8773 0.380324
## nodcat3 -0.012021 0.118491 -0.1014 0.919195
## nodcat4 0.143596 0.182086 0.7886 0.430336
## nodcat5-9 -0.482427 0.156551 -3.0816 0.002059 **
## nodcat10+ 0.043502 0.299620 0.1452 0.884561
## mets1 0.100543 0.099312 1.0124 0.311349
## mets2 0.139866 0.105583 1.3247 0.185272
## mets3 -0.091375 0.296308 -0.3084 0.757795
## durcat10-19m 0.038115 0.342352 0.1113 0.911352
## durcat20-29m -0.009245 0.341993 -0.0270 0.978434
## durcat30-39m 0.127019 0.356684 0.3561 0.721758
## durcat40+m 0.159252 0.394089 0.4041 0.686138
## diacat1.5-1.9cm -1.053938 0.100734 -10.4626 < 2.2e-16 ***
## diacat2-2.4cm -2.223150 0.109874 -20.2337 < 2.2e-16 ***
## diacat2.5-2.9cm -3.781319 0.199920 -18.9142 < 2.2e-16 ***
## diacat3cm+ -4.216794 0.470139 -8.9693 < 2.2e-16 ***
## primarybreast 0.070017 0.288955 0.2423 0.808540
## primarycolorectal 0.087730 0.292264 0.3002 0.764044
## primarygullet -0.091053 0.378701 -0.2404 0.809993
## primarykidney -0.269296 0.377865 -0.7127 0.476046
## primaryprostate -0.027718 0.344383 -0.0805 0.935850
## primaryskin 1.557764 0.670168 2.3244 0.020102 *
## primarystomach 0.134603 0.380885 0.3534 0.723792
## primarytesticular 0.304981 0.403875 0.7551 0.450167
## positionmoderate 0.891048 0.099587 8.9474 < 2.2e-16 ***
## positiondifficult 1.178730 0.116364 10.1296 < 2.2e-16 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## (Dispersion parameter for binomial family taken to be 1)
##
## Null deviance: 4916.81 on 3550 degrees of freedom
## Residual deviance: 3852.35 on 3513 degrees of freedom
## AIC: 3928.35
##
## Number of Fisher Scoring iterations: 5RFAcat$Score <- Pros_Score$fitted.values # extract the fitted scores
ggthemr('fresh', layout = 'scientific')
RFAcat %>%
ggplot(aes(x = Score, y= ..density.., fill = as.factor(rfa))) +
geom_histogram(position = "identity", color = "black", alpha = 0.5) +
geom_density(alpha = 0.2) +
theme(axis.title = element_text(size = 17), axis.text = element_text(size = 14),
axis.line = element_line(colour = "black"),
panel.border = element_blank(),
panel.background = element_blank()) +
theme(legend.text = element_text(size = 19),
legend.title = element_text(size = 19),
legend.position = "bottom", legend.direction = "horizontal") +
labs(fill = "Treatment Methods") +
scale_fill_discrete(labels = c("Standard surgery", "RFA")); ggthemr_reset()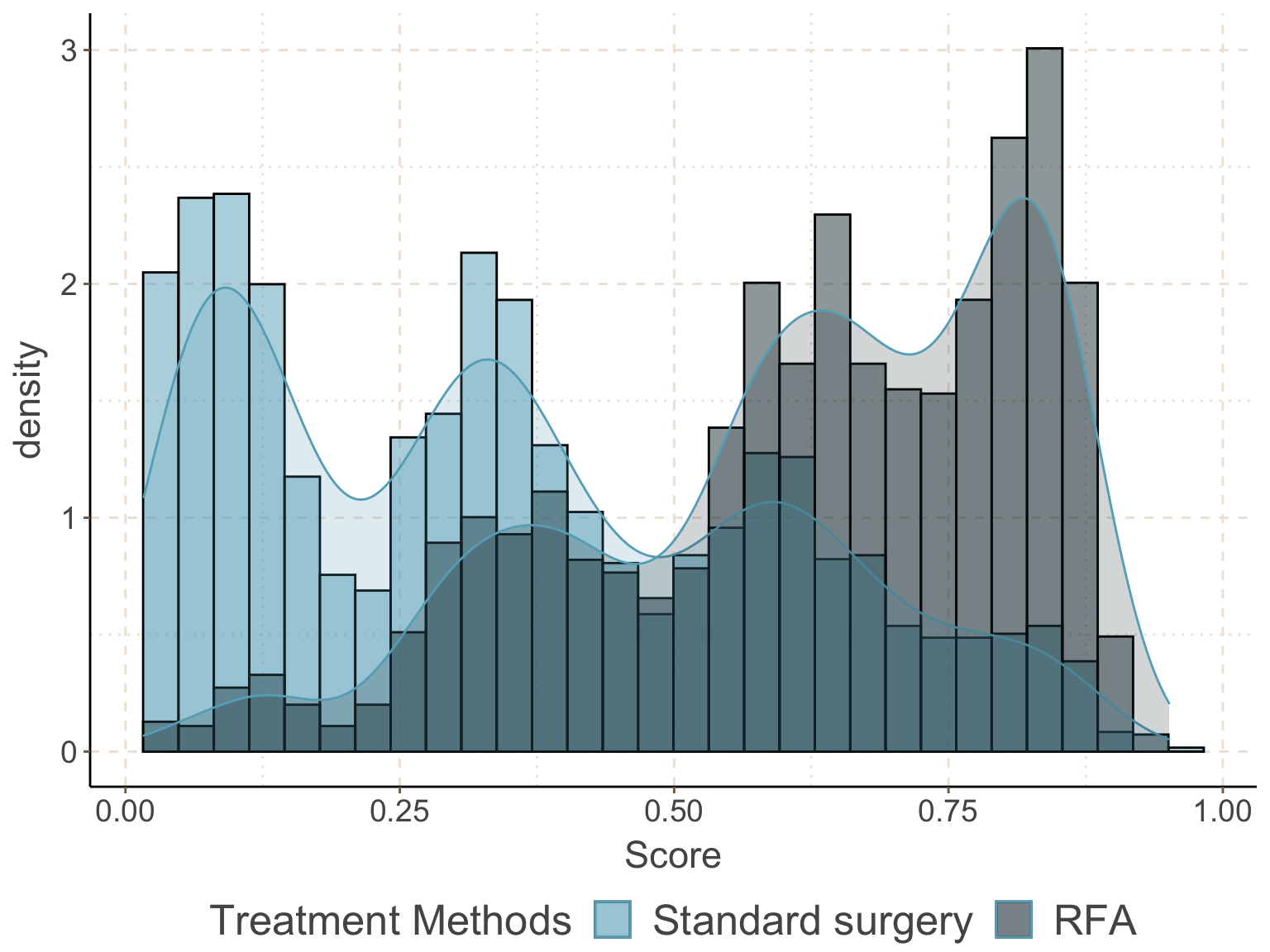
圖 111.1: Density and histogram of the estimated propensity score in the two exposure groups, with confounders and predictors of outcome included in the PS model.
概率密度分布圖和直方圖的內容告訴我們兩個暴露組患者的評分分布的交叉部分十分令人滿意,positivity 的前提假設可認爲得到滿足 (每個患者都有非零的概率接受 RFA 或者標準手術療法)。
一種比較組與組之間不同量的指標: standardized (mean) difference (Austin 2011),可以用下面的方法來計算,使用 tableone 這個方便的 R 包:
# covariates
Cov <- c("agecat", "gender", "smoke", "hospital", "nodcat", "mets", "durcat", "diacat",
"primary", "position")
## Construct a table
tabUnmatched <- CreateTableOne(vars = Cov, strata = "rfa", data = RFAcat, test = FALSE)
## Show table with SMD
print(tabUnmatched, smd = TRUE)## Stratified by rfa
## 0 1 SMD
## n 1848 1703
## agecat (%) 0.065
## < 45 51 ( 2.8) 45 ( 2.6)
## 45-49 463 (25.1) 397 (23.3)
## 50-54 891 (48.2) 864 (50.7)
## 55-59 352 (19.0) 310 (18.2)
## 60-64 72 ( 3.9) 63 ( 3.7)
## 65+ 19 ( 1.0) 24 ( 1.4)
## gender = women (%) 726 (39.3) 701 (41.2) 0.038
## smoke (%) 0.031
## never 1279 (69.2) 1157 (67.9)
## ex 135 ( 7.3) 136 ( 8.0)
## current 434 (23.5) 410 (24.1)
## hospital (%) 0.115
## 1 308 (16.7) 262 (15.4)
## 2 349 (18.9) 282 (16.6)
## 3 677 (36.6) 718 (42.2)
## 4 514 (27.8) 441 (25.9)
## nodcat (%) 0.096
## 1 505 (27.3) 492 (28.9)
## 2 703 (38.0) 647 (38.0)
## 3 325 (17.6) 310 (18.2)
## 4 104 ( 5.6) 100 ( 5.9)
## 5-9 181 ( 9.8) 123 ( 7.2)
## 10+ 30 ( 1.6) 31 ( 1.8)
## mets (%) 0.061
## 0 1089 (58.9) 969 (56.9)
## 1 393 (21.3) 375 (22.0)
## 2 327 (17.7) 331 (19.4)
## 3 39 ( 2.1) 28 ( 1.6)
## durcat (%) 0.070
## < 10m 31 ( 1.7) 22 ( 1.3)
## 10-19m 731 (39.6) 706 (41.5)
## 20-29m 820 (44.4) 708 (41.6)
## 30-39m 197 (10.7) 200 (11.7)
## 40+m 69 ( 3.7) 67 ( 3.9)
## diacat (%) 1.086
## <1.5cm 222 (12.0) 745 (43.7)
## 1.5-1.9cm 507 (27.4) 640 (37.6)
## 2-2.4cm 661 (35.8) 279 (16.4)
## 2.5-2.9cm 369 (20.0) 34 ( 2.0)
## 3cm+ 89 ( 4.8) 5 ( 0.3)
## primary (%) 0.126
## bladder 46 ( 2.5) 35 ( 2.1)
## breast 423 (22.9) 401 (23.5)
## colorectal 694 (37.6) 671 (39.4)
## gullet 75 ( 4.1) 55 ( 3.2)
## kidney 66 ( 3.6) 47 ( 2.8)
## prostate 365 (19.8) 324 (19.0)
## skin 5 ( 0.3) 17 ( 1.0)
## stomach 86 ( 4.7) 67 ( 3.9)
## testicular 88 ( 4.8) 86 ( 5.0)
## position (%) 0.332
## easy 565 (30.6) 292 (17.1)
## moderate 899 (48.6) 921 (54.1)
## difficult 384 (20.8) 490 (28.8)##
## FALSE TRUE Sum
## 5 5 10嚴重不平衡的變量有 5 個: hospital, nodcat, diaact, primary, position。
111.1.4 用 PS 評分來把對象分層 stratification
RFAcat <- RFAcat %>%
mutate(psblock = ntile(Score, 4))
RFAcat %>%
group_by(psblock) %>%
summarise(n(), min(Score), max(Score))## # A tibble: 4 × 4
## psblock `n()` `min(Score)` `max(Score)`
## <int> <int> <dbl> <dbl>
## 1 1 888 0.0167 0.280
## 2 2 888 0.281 0.508
## 3 3 888 0.508 0.695
## 4 4 887 0.696 0.951但是你看每層的傾向性評分其實範圍有點寬,提示使用分層的方法可能殘餘的混雜有點多。
看每層內數據的平衡:
# Cov <- c("diacat", "position")
#---------------------------------------------#
# in strata == 1 #
# #
#---------------------------------------------#
## Construct a table
tabUnmatched <- CreateTableOne(vars = Cov, strata = "rfa", data = RFAcat[RFAcat$psblock == 1,], test = FALSE)
## Show table with SMD
print(tabUnmatched, smd = TRUE)## Stratified by rfa
## 0 1 SMD
## n 777 111
## agecat (%) 0.321
## < 45 27 ( 3.5) 3 ( 2.7)
## 45-49 211 (27.2) 25 (22.5)
## 50-54 334 (43.0) 59 (53.2)
## 55-59 162 (20.8) 23 (20.7)
## 60-64 37 ( 4.8) 1 ( 0.9)
## 65+ 6 ( 0.8) 0 ( 0.0)
## gender = women (%) 304 (39.1) 45 (40.5) 0.029
## smoke (%) 0.248
## never 533 (68.6) 88 (79.3)
## ex 56 ( 7.2) 6 ( 5.4)
## current 188 (24.2) 17 (15.3)
## hospital (%) 0.265
## 1 135 (17.4) 10 ( 9.0)
## 2 152 (19.6) 27 (24.3)
## 3 270 (34.7) 38 (34.2)
## 4 220 (28.3) 36 (32.4)
## nodcat (%) 0.209
## 1 201 (25.9) 24 (21.6)
## 2 293 (37.7) 47 (42.3)
## 3 119 (15.3) 17 (15.3)
## 4 41 ( 5.3) 7 ( 6.3)
## 5-9 112 (14.4) 16 (14.4)
## 10+ 11 ( 1.4) 0 ( 0.0)
## mets (%) 0.141
## 0 463 (59.6) 70 (63.1)
## 1 154 (19.8) 24 (21.6)
## 2 138 (17.8) 15 (13.5)
## 3 22 ( 2.8) 2 ( 1.8)
## durcat (%) 0.209
## < 10m 17 ( 2.2) 3 ( 2.7)
## 10-19m 297 (38.2) 41 (36.9)
## 20-29m 356 (45.8) 54 (48.6)
## 30-39m 76 ( 9.8) 12 (10.8)
## 40+m 31 ( 4.0) 1 ( 0.9)
## diacat (%) 0.515
## <1.5cm 0 ( 0.0) 0 ( 0.0)
## 1.5-1.9cm 9 ( 1.2) 4 ( 3.6)
## 2-2.4cm 310 (39.9) 68 (61.3)
## 2.5-2.9cm 369 (47.5) 34 (30.6)
## 3cm+ 89 (11.5) 5 ( 4.5)
## primary (%) 0.279
## bladder 23 ( 3.0) 1 ( 0.9)
## breast 180 (23.2) 25 (22.5)
## colorectal 273 (35.1) 43 (38.7)
## gullet 40 ( 5.1) 3 ( 2.7)
## kidney 34 ( 4.4) 7 ( 6.3)
## prostate 159 (20.5) 25 (22.5)
## skin 1 ( 0.1) 1 ( 0.9)
## stomach 32 ( 4.1) 3 ( 2.7)
## testicular 35 ( 4.5) 3 ( 2.7)
## position (%) 0.181
## easy 319 (41.1) 36 (32.4)
## moderate 342 (44.0) 55 (49.5)
## difficult 116 (14.9) 20 (18.0)##
## FALSE TRUE Sum
## 1 9 10#---------------------------------------------#
# in strata == 2 #
# #
#---------------------------------------------#
## Construct a table
tabUnmatched <- CreateTableOne(vars = Cov, strata = "rfa", data = RFAcat[RFAcat$psblock == 2,], test = FALSE)
## Show table with SMD
print(tabUnmatched, smd = TRUE)## Stratified by rfa
## 0 1 SMD
## n 550 338
## agecat (%) 0.114
## < 45 12 ( 2.2) 5 ( 1.5)
## 45-49 129 (23.5) 82 (24.3)
## 50-54 294 (53.5) 173 (51.2)
## 55-59 87 (15.8) 61 (18.0)
## 60-64 21 ( 3.8) 10 ( 3.0)
## 65+ 7 ( 1.3) 7 ( 2.1)
## gender = women (%) 217 (39.5) 132 (39.1) 0.008
## smoke (%) 0.091
## never 384 (69.8) 222 (65.7)
## ex 39 ( 7.1) 29 ( 8.6)
## current 127 (23.1) 87 (25.7)
## hospital (%) 0.101
## 1 91 (16.5) 48 (14.2)
## 2 89 (16.2) 49 (14.5)
## 3 226 (41.1) 154 (45.6)
## 4 144 (26.2) 87 (25.7)
## nodcat (%) 0.139
## 1 150 (27.3) 102 (30.2)
## 2 205 (37.3) 124 (36.7)
## 3 117 (21.3) 61 (18.0)
## 4 37 ( 6.7) 18 ( 5.3)
## 5-9 30 ( 5.5) 26 ( 7.7)
## 10+ 11 ( 2.0) 7 ( 2.1)
## mets (%) 0.145
## 0 322 (58.5) 191 (56.5)
## 1 127 (23.1) 72 (21.3)
## 2 93 (16.9) 73 (21.6)
## 3 8 ( 1.5) 2 ( 0.6)
## durcat (%) 0.162
## < 10m 8 ( 1.5) 4 ( 1.2)
## 10-19m 210 (38.2) 138 (40.8)
## 20-29m 251 (45.6) 133 (39.3)
## 30-39m 63 (11.5) 44 (13.0)
## 40+m 18 ( 3.3) 19 ( 5.6)
## diacat (%) 0.057
## <1.5cm 4 ( 0.7) 4 ( 1.2)
## 1.5-1.9cm 199 (36.2) 127 (37.6)
## 2-2.4cm 347 (63.1) 207 (61.2)
## 2.5-2.9cm 0 ( 0.0) 0 ( 0.0)
## 3cm+ 0 ( 0.0) 0 ( 0.0)
## primary (%) 0.099
## bladder 9 ( 1.6) 8 ( 2.4)
## breast 124 (22.5) 79 (23.4)
## colorectal 221 (40.2) 126 (37.3)
## gullet 19 ( 3.5) 13 ( 3.8)
## kidney 14 ( 2.5) 12 ( 3.6)
## prostate 104 (18.9) 63 (18.6)
## skin 0 ( 0.0) 0 ( 0.0)
## stomach 33 ( 6.0) 19 ( 5.6)
## testicular 26 ( 4.7) 18 ( 5.3)
## position (%) 0.041
## easy 172 (31.3) 100 (29.6)
## moderate 242 (44.0) 155 (45.9)
## difficult 136 (24.7) 83 (24.6)##
## FALSE TRUE Sum
## 3 7 10#---------------------------------------------#
# in strata == 3 #
# #
#---------------------------------------------#
## Construct a table
tabUnmatched <- CreateTableOne(vars = Cov, strata = "rfa", data = RFAcat[RFAcat$psblock == 3,], test = FALSE)
## Show table with SMD
print(tabUnmatched, smd = TRUE)## Stratified by rfa
## 0 1 SMD
## n 347 541
## agecat (%) 0.163
## < 45 6 ( 1.7) 13 ( 2.4)
## 45-49 90 (25.9) 135 (25.0)
## 50-54 171 (49.3) 261 (48.2)
## 55-59 71 (20.5) 102 (18.9)
## 60-64 8 ( 2.3) 25 ( 4.6)
## 65+ 1 ( 0.3) 5 ( 0.9)
## gender = women (%) 132 (38.0) 205 (37.9) 0.003
## smoke (%) 0.049
## never 245 (70.6) 372 (68.8)
## ex 22 ( 6.3) 40 ( 7.4)
## current 80 (23.1) 129 (23.8)
## hospital (%) 0.125
## 1 59 (17.0) 107 (19.8)
## 2 78 (22.5) 109 (20.1)
## 3 101 (29.1) 176 (32.5)
## 4 109 (31.4) 149 (27.5)
## nodcat (%) 0.087
## 1 101 (29.1) 161 (29.8)
## 2 143 (41.2) 205 (37.9)
## 3 59 (17.0) 101 (18.7)
## 4 14 ( 4.0) 24 ( 4.4)
## 5-9 22 ( 6.3) 40 ( 7.4)
## 10+ 8 ( 2.3) 10 ( 1.8)
## mets (%) 0.061
## 0 205 (59.1) 314 (58.0)
## 1 78 (22.5) 122 (22.6)
## 2 57 (16.4) 89 (16.5)
## 3 7 ( 2.0) 16 ( 3.0)
## durcat (%) 0.088
## < 10m 3 ( 0.9) 6 ( 1.1)
## 10-19m 150 (43.2) 227 (42.0)
## 20-29m 142 (40.9) 239 (44.2)
## 30-39m 40 (11.5) 51 ( 9.4)
## 40+m 12 ( 3.5) 18 ( 3.3)
## diacat (%) 0.142
## <1.5cm 68 (19.6) 135 (25.0)
## 1.5-1.9cm 275 (79.3) 403 (74.5)
## 2-2.4cm 4 ( 1.2) 3 ( 0.6)
## 2.5-2.9cm 0 ( 0.0) 0 ( 0.0)
## 3cm+ 0 ( 0.0) 0 ( 0.0)
## primary (%) 0.121
## bladder 12 ( 3.5) 18 ( 3.3)
## breast 73 (21.0) 124 (22.9)
## colorectal 131 (37.8) 198 (36.6)
## gullet 10 ( 2.9) 19 ( 3.5)
## kidney 13 ( 3.7) 16 ( 3.0)
## prostate 68 (19.6) 106 (19.6)
## skin 3 ( 0.9) 1 ( 0.2)
## stomach 17 ( 4.9) 25 ( 4.6)
## testicular 20 ( 5.8) 34 ( 6.3)
## position (%) 0.162
## easy 65 (18.7) 127 (23.5)
## moderate 216 (62.2) 294 (54.3)
## difficult 66 (19.0) 120 (22.2)##
## FALSE TRUE Sum
## 3 7 10#---------------------------------------------#
# in strata == 4 #
# #
#---------------------------------------------#
## Construct a table
tabUnmatched <- CreateTableOne(vars = Cov, strata = "rfa", data = RFAcat[RFAcat$psblock == 4,], test = FALSE)
## Show table with SMD
print(tabUnmatched, smd = TRUE)## Stratified by rfa
## 0 1 SMD
## n 174 713
## agecat (%) 0.105
## < 45 6 ( 3.4) 24 ( 3.4)
## 45-49 33 (19.0) 155 (21.7)
## 50-54 92 (52.9) 371 (52.0)
## 55-59 32 (18.4) 124 (17.4)
## 60-64 6 ( 3.4) 27 ( 3.8)
## 65+ 5 ( 2.9) 12 ( 1.7)
## gender = women (%) 73 (42.0) 319 (44.7) 0.056
## smoke (%) 0.077
## never 117 (67.2) 475 (66.6)
## ex 18 (10.3) 61 ( 8.6)
## current 39 (22.4) 177 (24.8)
## hospital (%) 0.104
## 1 23 (13.2) 97 (13.6)
## 2 30 (17.2) 97 (13.6)
## 3 80 (46.0) 350 (49.1)
## 4 41 (23.6) 169 (23.7)
## nodcat (%) 0.254
## 1 53 (30.5) 205 (28.8)
## 2 62 (35.6) 271 (38.0)
## 3 30 (17.2) 131 (18.4)
## 4 12 ( 6.9) 51 ( 7.2)
## 5-9 17 ( 9.8) 41 ( 5.8)
## 10+ 0 ( 0.0) 14 ( 2.0)
## mets (%) 0.061
## 0 99 (56.9) 394 (55.3)
## 1 34 (19.5) 157 (22.0)
## 2 39 (22.4) 154 (21.6)
## 3 2 ( 1.1) 8 ( 1.1)
## durcat (%) 0.094
## < 10m 3 ( 1.7) 9 ( 1.3)
## 10-19m 74 (42.5) 300 (42.1)
## 20-29m 71 (40.8) 282 (39.6)
## 30-39m 18 (10.3) 93 (13.0)
## 40+m 8 ( 4.6) 29 ( 4.1)
## diacat (%) 0.062
## <1.5cm 150 (86.2) 606 (85.0)
## 1.5-1.9cm 24 (13.8) 106 (14.9)
## 2-2.4cm 0 ( 0.0) 1 ( 0.1)
## 2.5-2.9cm 0 ( 0.0) 0 ( 0.0)
## 3cm+ 0 ( 0.0) 0 ( 0.0)
## primary (%) 0.177
## bladder 2 ( 1.1) 8 ( 1.1)
## breast 46 (26.4) 173 (24.3)
## colorectal 69 (39.7) 304 (42.6)
## gullet 6 ( 3.4) 20 ( 2.8)
## kidney 5 ( 2.9) 12 ( 1.7)
## prostate 34 (19.5) 130 (18.2)
## skin 1 ( 0.6) 15 ( 2.1)
## stomach 4 ( 2.3) 20 ( 2.8)
## testicular 7 ( 4.0) 31 ( 4.3)
## position (%) 0.056
## easy 9 ( 5.2) 29 ( 4.1)
## moderate 99 (56.9) 417 (58.5)
## difficult 66 (37.9) 267 (37.4)##
## FALSE TRUE Sum
## 5 5 10可以看出其實分層法中每層的數據依然還有很多的不平衡。分層法不是合理的利用傾向性評分的理想辦法。
111.1.4.1 計算每層評分組內,暴露和結果之間的關系
##
## Call:
## lm(formula = dodp ~ rfa, data = RFAcat[RFAcat$psblock == 1, ])
##
## Coefficients:
## (Intercept) rfa
## 0.40927 0.11326##
## Call:
## lm(formula = dodp ~ rfa, data = RFAcat[RFAcat$psblock == 2, ])
##
## Coefficients:
## (Intercept) rfa
## 0.285455 0.099161##
## Call:
## lm(formula = dodp ~ rfa, data = RFAcat[RFAcat$psblock == 3, ])
##
## Coefficients:
## (Intercept) rfa
## 0.198847 -0.041731##
## Call:
## lm(formula = dodp ~ rfa, data = RFAcat[RFAcat$psblock == 4, ])
##
## Coefficients:
## (Intercept) rfa
## 0.28161 -0.16800111.1.5 用配對法計算 ACE
##
## . use "../backupfiles/RFAcat.dt. * Question 10
##
## . teffects psmatch (dodp) (rfa i.agecat gender i.smoke i.hospital i.nodcat i.me
## > ts i.durcat i.diacat i.primary i.position, logit)
##
## Treatment-effects estimation Number of obs = 3,551
## Estimator : propensity-score matching Matches: requested = 1
## Outcome model : matching min = 1
## Treatment model: logit max = 3
## ------------------------------------------------------------------------------
## | AI Robust
## dodp | Coef. Std. Err. z P>|z| [95% Conf. Interval]
## -------------+----------------------------------------------------------------
## ATE |
## rfa |
## (radiofre.. |
## vs |
## standard..) | .0263775 .0187999 1.40 0.161 -.0104697 .0632248
## ------------------------------------------------------------------------------
##
## .111.1.6 模型校正 PS
RFAcat$rfanew <- RFAcat$rfa
Log_ps <- glm(dodp ~ as.factor(rfanew)*Score, data = RFAcat, family = binomial(link = "logit"))
summary(Log_ps)##
## Call:
## glm(formula = dodp ~ as.factor(rfanew) * Score, family = binomial(link = "logit"),
## data = RFAcat)
##
## Deviance Residuals:
## Min 1Q Median 3Q Max
## -1.45859 -0.86831 -0.63218 1.29348 2.29729
##
## Coefficients:
## Estimate Std. Error z value Pr(>|z|)
## (Intercept) -0.28539 0.08742 -3.2646 0.001096 **
## as.factor(rfanew)1 1.04607 0.19586 5.3409 9.247e-08 ***
## Score -1.37952 0.22182 -6.2190 5.003e-10 ***
## as.factor(rfanew)1:Score -2.24706 0.37406 -6.0073 1.886e-09 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## (Dispersion parameter for binomial family taken to be 1)
##
## Null deviance: 4118.69 on 3550 degrees of freedom
## Residual deviance: 3863.82 on 3547 degrees of freedom
## AIC: 3871.82
##
## Number of Fisher Scoring iterations: 4RFAcat <- RFAcat %>%
mutate(rfanew = 1)
newdata <- subset(RFAcat, select = c(rfanew, Score))
RFAcat$Po1<- predict(Log_ps, newdata, type = "response")
RFAcat <- RFAcat %>%
mutate(rfanew = 0)
newdata1 <- subset(RFAcat, select = c(rfanew, Score))
RFAcat$Po0 <- predict(Log_ps, newdata1, type = "response")
with(RFAcat, epiDisplay::summ(Po1, graph = F))## obs. mean median s.d. min. max.
## 3551 0.306 0.253 0.182 0.064 0.668## obs. mean median s.d. min. max.
## 3551 0.285 0.272 0.072 0.168 0.423## obs. mean median s.d. min. max.
## 3551 0.021 -0.018 0.11 -0.105 0.245References
Austin, Peter C. 2011. “An Introduction to Propensity Score Methods for Reducing the Effects of Confounding in Observational Studies.” Multivariate Behavioral Research 46 (3): 399–424.